P: Mass calibration (single-dataset)
Brief description
Calibrate mass axis.
Parameters
m/zs (type - array)
M/z values.
m/zs (must be used during calibration) (type - array)
M/z values that must be selected in the RANSAC process - select m/zs that will ensure convergence of the calibration procedure.
Help
We are using RANSAC(RANdom SAmple Consensus) algorithm to improve the mass calibration performance
which randomly samples the data and fits a model to the data. This is particularly useful when the data
is noisy or contains outliers. By specifying the essential m/zs, you can ensure that certain mass
ranges are always included in the calibration process.
Method (type - string)
Method to use for calibration. 'regression' is much better than 'interpolation'
More Information
The mass calibration task is a critical step in the pre-processing of mass spectrometry imaging data. It ensures the accuracy and precision of mass measurements, which is essential for downstream analysis.
The mass alignment task task removed majority of misalignment in the dataset (shifts along the mass axis due to change of temperature or other factors), but it would not correct for any miscalibration.
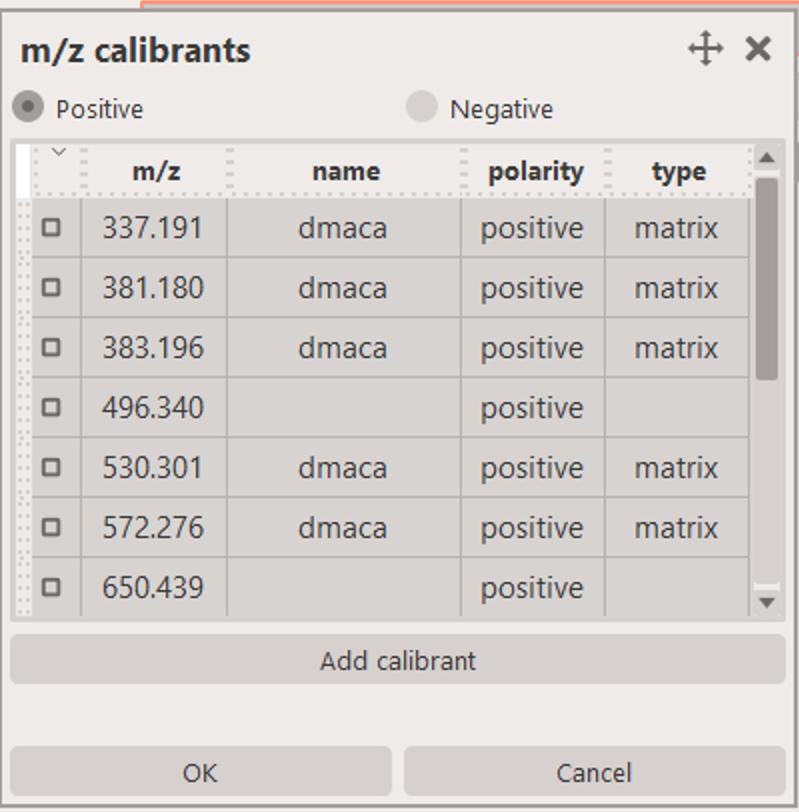
This task relies on you specifing several (at least 2-3) calibrant species by either typing them in (e.g. 496.340,650.439,703.575,734.569,760.585) or selecting them from the list of available calibrants.
Outputs
The mass calibration task generates a couple of outputs that can help diagnose how well the task went and also the quality of your data.
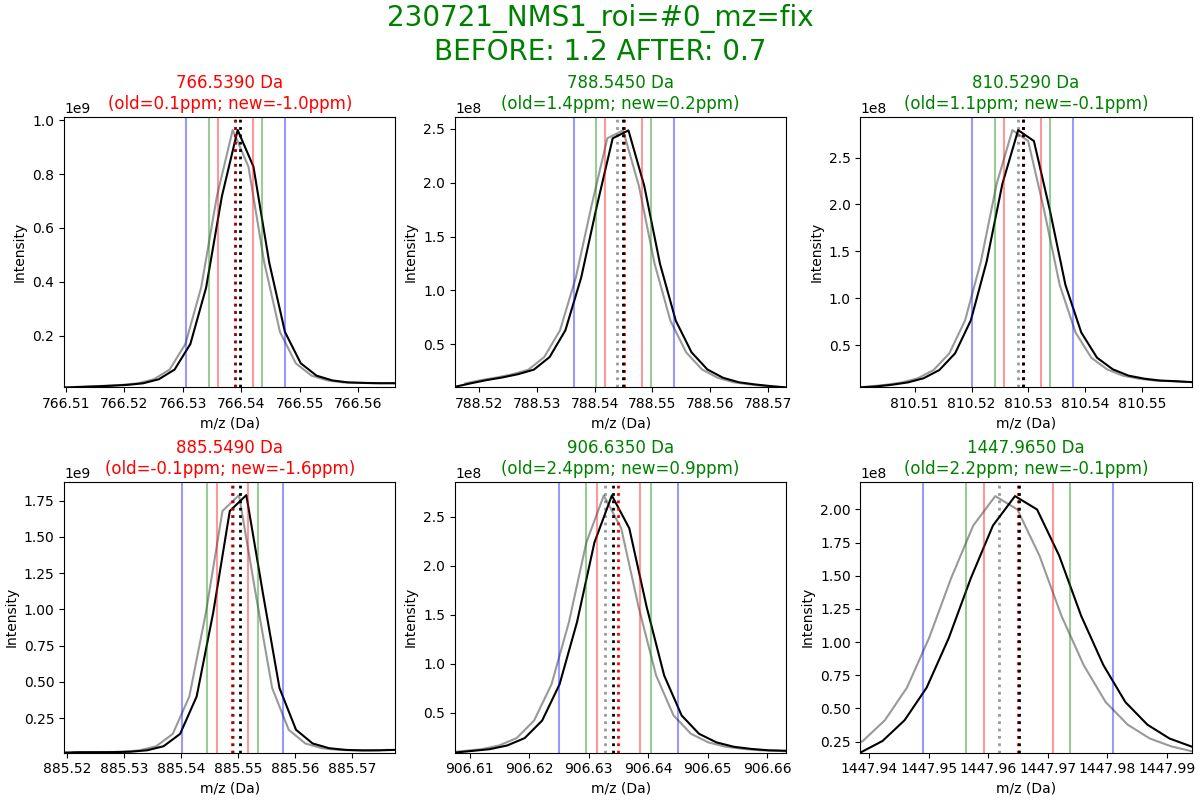
Before and after mass calibration
This plot shows you the effect the calibration had on your data. It will show a peak before and after calibration and you can see how much the mass axis shifted.
Thigs of note
- one file will be generated for each dataset (you can see the dataset name in the title of the plot)
- the title will show you the ppm error before and after calibration
- the plot will show you the mass axis before and after calibration
- the before and after mass error is also shown for each peak (m/zs selected for calibration)
- values in green indicate that ppm error was reduced, and values in red indicate that ppm error was increased
Dependencies (other tasks that this task might depend on)
| Depends on | Required/Optional |
|---|---|
| P: Average mass spectrum (single-dataset) | required |
| W: Average mass spectrum / mask (single-dataset) | required |
| W: Average mass spectrum / cluster (single-dataset) | required |
Attributes
| Attribute | Value | Description |
|---|---|---|
| Multiple allowed | False | Allow multiple instances of this task in a workflow. |
| Task can fail | False | Task is optional and can fail without causing the entire workflow to fail. |
| Step can fail | True | Sub-tasks of this task can fail without causing the entire task (and workflow) to fail. |
| Requires ion mobility | False | Task requires ion mobility data. |
| Task can fail (with ion mobility) | False | This task uses ion mobility data but it is allowed to fail, without causing the entire workflow to fail. |
| Allowed in reference dataset | False | Task is to be performed on a 'reference' dataset. This will allow for multiple analyses to be performed on the same dataset, without cluttering or duplicating certain tasks (unused at the moment). |